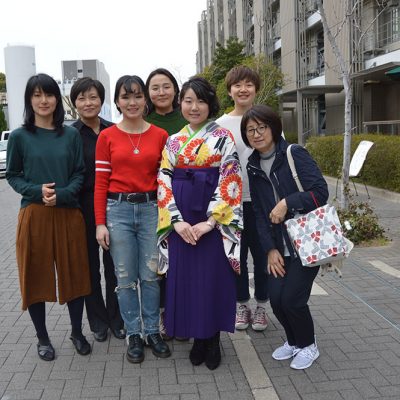
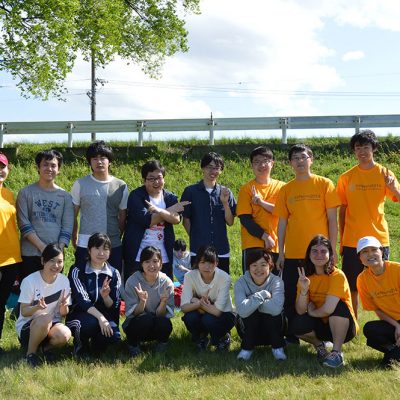
You Lab
We are interested in how metabolism regulates various behaviors. We use two invertebrate model systems of C. elegans and D. melanogaster, ultimately aiming to unveil conserved neuro-molecular mechanisms throughout animals including mammals.
Find us on Twitter @ZigmanLab
Read About us in the news

Some of our major ongoing projects are listed here:
Neural mechanism of starvation
Metabolic regulation of reproduction
Metabolic regulation of behavioral states

Neural Mechanism of Starvation
Long-term nutrient deprivation associates with dramatic changes in development and behavior. However, how nutrient deprivation induces particular neural pathways to control behavioral and metabolic adaptations is not fully understood. We previously found that the feeding organ of C. elegans undergoes changes to adapt starvation (You et al, 2006) via a muscarinic receptor to the MAPK pathway, identifying a role of acetylcholine as a part of a starvation signaling pathway. Based on the finding, we aim to further identify neuronal starvation signals.

Metabolic Regulation of Reproduction
In the wild, animals often face a serious challenge to maintain oocyte quiescence under long-lasting unfavorable conditions in the absence of mates or food. Failure to maintain oocyte quiescence will result in activation of oocytes to maturation at the wrong time, and thus lead to exhaustion of the oocyte pool and sterility of the organism. Current pandemic dysregulation of metabolism manifests the connection between metabolism and reproduction, shown in polycystic ovary syndrome (PCOS). Using our in-depth knowledge in starvation and metabolism, we aim to investigate how metabolism regulates oocyte arrest and maturation using C. elegans and D. melanogaster, in collaboration with Dr. Jeongho Kim in Inha University of Korea.

Metabolic Regulation of Behavioral States
Animals change their feeding behavior based on the past experiences and nutritional or developmental needs. Hungry animals leave safety to seek food. Satiated animals stop seeking food and often sleep. Animals control feeding behavior by integrating multiple internal and external cues. Misregulation of this process causes disorders from anorexia to obesity. Previously we found that C. elegans exhibits conserved hunger and satiety behaviors. We study the feeding circuitry and molecular mechanisms by which multiple cues are integrated and the proper feeding behavior is chosen, using C. elegans. One particular output reflecting satiety is sleep. Interestingly, circadian clock, a main controller of sleep timing, can be entrained by feeding. In addition, orexin, a peptide hormone, regulates both sleep and feeding, suggesting feeding and sleep are closely linked. Therefore, we are also investigating the molecular and neuronal mechanisms by which feeding and metabolism regulates sleep.

Metabolic Regulation of Behavioral States
Animals change their feeding behavior based on the past experiences and nutritional or developmental needs. Hungry animals leave safety to seek food. Satiated animals stop seeking food and often sleep. Animals control feeding behavior by integrating multiple internal and external cues. Misregulation of this process causes disorders from anorexia to obesity. Previously we found that C. elegans exhibits conserved hunger and satiety behaviors. We study the feeding circuitry and molecular mechanisms by which multiple cues are integrated and the proper feeding behavior is chosen, using C. elegans. One particular output reflecting satiety is sleep. Interestingly, circadian clock, a main controller of sleep timing, can be entrained by feeding. In addition, orexin, a peptide hormone, regulates both sleep and feeding, suggesting feeding and sleep are closely linked. Therefore, we are also investigating the molecular and neuronal mechanisms by which feeding and metabolism regulates sleep.

Neural Mechanism of Starvation
Long-term nutrient deprivation associates with dramatic changes in development and behavior. However, how nutrient deprivation induces particular neural pathways to control behavioral and metabolic adaptations is not fully understood. We previously found that the feeding organ of C. elegans undergoes changes to adapt starvation (You et al, 2006) via a muscarinic receptor to the MAPK pathway, identifying a role of acetylcholine as a part of a starvation signaling pathway. Based on the finding, we aim to further identify neuronal starvation signals.

Metabolic Regulation of Reproduction
In the wild, animals often face a serious challenge to maintain oocyte quiescence under long-lasting unfavorable conditions in the absence of mates or food. Failure to maintain oocyte quiescence will result in activation of oocytes to maturation at the wrong time, and thus lead to exhaustion of the oocyte pool and sterility of the organism. Current pandemic dysregulation of metabolism manifests the connection between metabolism and reproduction, shown in polycystic ovary syndrome (PCOS). Using our in-depth knowledge in starvation and metabolism, we aim to investigate how metabolism regulates oocyte arrest and maturation using C. elegans and D. melanogaster, in collaboration with Dr. Jeongho Kim in Inha University of Korea.

Metabolic Regulation of Behavioral States
Animals change their feeding behavior based on the past experiences and nutritional or developmental needs. Hungry animals leave safety to seek food. Satiated animals stop seeking food and often sleep (Movie 1 & 2). Animals control feeding behavior by integrating multiple internal and external cues. Misregulation of this process causes disorders from anorexia to obesity. Previously we found that C. elegans exhibits conserved hunger and satiety behaviors. We study the feeding circuitry and molecular mechanisms by which multiple cues are integrated and the proper feeding behavior is chosen, using C. elegans. One particular output reflecting satiety is sleep. Interestingly, circadian clock, a main controller of sleep timing, can be entrained by feeding. In addition, orexin, a peptide hormone, regulates both sleep and feeding, suggesting feeding and sleep are closely linked. Therefore, we are also investigating the molecular and neuronal mechanisms by which feeding and metabolism regulates sleep (Fig. 2).
You Lab
Details about Our Research
The Zigman Lab is composed of Dr. Zigman, our lab manager and senior research scientist, Sherri Osborne-Lawrence, and a core group of talented postdoctoral fellows/students/research associates/technicians. We investigate how gastrointestinal hormones such as ghrelin and LEAP2 interact with the brain and peripheral nervous system to control eating, body weight, blood glucose, responses to exercise, and mood in an effort to devise better treatments for obesity, diabetes, eating disorders, and depression.
Our work spans the fields of molecular endocrinology, behavioral neuroscience, and neuroanatomy and includes mouse, human, and cell culture experiments.
Our studies employ a one-of-a-kind collection of Zigman Lab-generated novel transgenic mouse models including those made using recombineering and CRISPR-Cas9 approaches. We also design and use several animal behavioral models and cell culture models. We incorporate state-of-the-art techniques to characterize gene expression, manipulate gene expression, and modulate cellular activity in specific cells and neurons of interest, for instance by chemogenetic/DREADD technology, optogenetics, and in vivo fiber photometry, to name a few. We use gold standard methodology such as glucose clamps to investigate glucose homeostasis. We also routinely utilize RNAseq technology, tissue clearing, cell culture, traditional neuroanatomic techniques, microscopy of various types, and histochemistry to probe ghrelin action, ghrelin secretion, and the coordinated hormonal/neuronal response to conditions relevant to perturbed metabolic states.
Milestones
We were first to define essential roles for ghrelin in mediating stress-induced comfort food eating and other complex eating behaviors, including operant responding and conditioned place preference for high-fat diet, also confirming a role for the ghrelin system in cue-potentiated feeding. We also have characterized the role of ghrelin in post-exercise eating.
We have contributed substantively to a body of work demonstrating an essential role for the ghrelin system in the regulation of blood glucose. We have demonstrated not only that ghrelin secretion is enhanced when ghrelin cells are exposed to low glucose, but also that when regulation of ghrelin secretion by the sympathetic nervous system is blocked, life-threatening falls in blood glucose ensue. We have also identified the arcuate nucleus, the caudal brainstem, and pancreatic alpha cells as targets for ghrelin’s glucoregulatory actions.
Consistent with our mission to understand the biological basis for the strong link between eating behavior and mood, our group was the first to demonstrate a role for the hormone ghrelin as a natural antidepressant, preventing exaggerated depressive-like behaviors following chronic stress and inducing an antidepressant-like behavioral response upon caloric restriction. We showed that ghrelin’s antidepressant actions rely on protection of adult hippocampal neurogenesis, leading us in collaborative work with Andrew Pieper to identify antidepressant-like efficacy for the P7C3 class of strong, rapid-acting neuroprotective compounds, and paving the way
We have identified key central and peripheral sites of ghrelin’s orexigenic, glucoregulatory and antidepressant actions by comprehensively determining the pattern of ghrelin receptor-expressing neurons in the rat and mouse brain, by using Cre-lox mouse genetics to target ghrelin receptor expression to selective cell-types, and by using chemogenetics to manipulate the activity of ghrelin receptor-expressing neurons in specific brain regions.
Using a collection of novel models to directly study isolated populations of ghrelin cells and to modify ghrelin cell gene expression, we have led the way in identifying direct modulators of ghrelin release and key elements of the ghrelin cell molecular machinery mediating ghrelin secretion.
We determined that circulating levels of the endogenous ghrelin receptor antagonist and inverse agonist LEAP-2 in humans are correlated with body weight, several metabolic parameters related to obesity, food intake, and weight loss.
We have generated a large toolbox of transgenic mouse models with which to investigate ghrelin action in the brain and periphery, ghrelin cell biology, and the biology of other gastric enteroendocrine cell types. These include ghrelin-Cre mice, GHSR-IRES-Cre mice, HDC-Cre mice, SF1-Cre mice, KISS1-Cre mice, “floxed”-β1AR mice, ghrelin-KO mice, GHSR-null mice, and ghrelin-hrGFP mice, to name a few. In collaboration with the Brown/Goldstein labs, we generated SG-1 and PG-1 immortalized ghrelin cell lines. These tools are now used in numerous metabolism and neuroscience research labs worldwide.
Current Lab Members

Sherri Osborne-Lawrence, M.S.
Senior Research Scientist and Lab Manager
Sherri Osborne-Lawrence, M.S.
Senior Research Scientist and Lab Manager
Sherri was born in Texas and has lived here all of her life. She received her B.A. degree in Biology from Austin College in Sherman, TX, and an M.S. degree in Molecular & Cell Biology from UT Dallas. She has worked in five laboratories at UT Southwestern Medical Center. Sherri’s initial research projects involved positional cloning of genes involved in rheumatoid arthritis and early-onset familial breast cancer. After a few years assisting with the final stages of sequencing for the Human Genome Project, she focused on discovering how endothelial cell dysfunction leads to cardiovascular disease, with particular interests in estrogen modulation as well as the effects of C-reactive protein. Sherri joined the Zigman Lab in 2006 as Senior Research Scientist and Lab Manager, and has now expanded her interests to include ghrelin and its effects on metabolic pathways. Among many other contributions, Sherri has designed and generated several new mouse lines currently in use in the lab and by our collaborators.

Nathan Metzger, M.S.
Research Associate
Nathan Metzger, M.S.
Research Associate
Nathan was born and raised in Apple Valley, MN. He obtained his BS in Community Health (2012) and Biomedical Science (2012) and his MS in Cell and Molecular Biology (2014) from Saint Cloud State University in Saint Cloud, MN. His research background includes having studied the post-translational modifications of a family of proteins collectively known as PGC-1. Specifically, identifying modifiers, locating binding sites, and exploring their effects on PGC-1 expression and activity. He relocated to Fort Worth, TX in the summer of 2016, after which he joined Dr. Zigman’s lab.

Deepali Gupta, Ph.D.
Postdoctoral Research Fellow
Deepali Gupta, Ph.D.
Postdoctoral Research Fellow
Deepali was born in Chhattisgarh, a tribal state in the central part of India. She received her Bachelor in Pharmacy from Rajiv Gandhi Technical University in 2008 and Master in Pharmacy from BITS-Pilani, Pilani Campus, India in 2011. During her Master’s dissertation program mentored by Prof. Mahesh, she studied the role of serotonin signaling and hypothalamic-pituitary-adrenal axis dysregulation in depression. Later, she enrolled in the doctoral program at BITS-Pilani and continued to work under the guidance of Prof. Mahesh . Her doctoral research project was aimed at investigating the role of serotonin type-3 (5-HT3) receptor signaling in the pathogenesis of type-1 diabetes-induced depression and the antidepressant activity of novel 5-HT3 receptor antagonists. Deepali joined the Zigman lab in January 2017 to further explore pathogenic pathways involved in depression, in particular, those modulated by the ghrelin system. When not in the lab, she likes cooking and chatting with her family and friends.

Kripa Shankar, Ph.D.
Postdoctoral Research Fellow
Kripa Shankar, Ph.D.
Postdoctoral Research Fellow
Kripa was born in a small city Kannauj, famous as a city of perfume, in Uttar Pradesh, India. Kripa completed his undergraduate education at CSJM University in Kanpur, India. Afterwards, Kripa received a Master of Science degree in Biotechnology at Babasaheb Bhimrao Ambedkar University in Lucknow, India. After obtaining Junior and Senior Research fellowships from the University Grant Commission, India, Kripa joined the lab of Dr. Anil N. Gaikwad at CSIR-Central Drug Research Institute in Lucknow, India. He earned his PhD degree in 2017. His PhD work focused on pathways involved in chronic hyperinsulinemia and high fat diet-induced insulin resistance. In June of 2017, Kripa joined the Zigman Lab to further enhance his expertise in the area of ghrelin-mediated effects on blood glucose regulation. Apart from science, Kripa loves watching cricket and listening to Bollywood music.

Salil Varshney, Ph.D.
Postdoctoral Research Fellow
Salil Varshney, Ph.D.
Postdoctoral Research Fellow
Salil was born and raised in Chandausi, a large town in Sambhal district, Uttar Pradesh, India. Salil completed his undergraduate education at M.J.P. Rohilkhand University, S.M. Degree College, Chandausi India. Afterwards, Salil received a Master of Science degree in Microbiology at University of Lucknow, India while working with Dr. Anil N. Gaikwad. Afterwards, Salil worked as a research fellow in India, where he was engaged in an institutional drug discovery program. After obtaining Senior Research fellowships from the Indian Council of Medical Research (ICMR) New Delhi, India, Salil enrolled in the doctoral program at CSIR- Central Drug Research Institute and continued to work with Dr. Gaikwad. He earned his PhD degree in 2019 from the Academy of Scientific and Innovative Research New Delhi, India. His PhD work focused on Harnessing good of lipids for healthy lifestyle: lipid intervention from signalling to phenotype in adipocyte biology and insulin resistance. After completing his doctoral training in the field of diabetes and obesity, Salil was interested in advanced training in diabetes and obesity. This led him to join the Zigman Lab in Feb 2020. Outside the lab, Salil loves playing & watching cricket, listening to music, and spending time with family and friends.

Sean Ogden, Ph.D.
Postdoctoral Research Fellow
Sean Ogden, Ph.D.
Postdoctoral Research Fellow
Sean was born and raised in San Diego, CA. In May 2011, Sean received his B.A. in Behavioral Neuroscience from Purdue University in West Lafayette, IN. While at Purdue, he trained under Drs. Susie Swithers and Terry Davidson investigating the effects of artificial sweeteners on food intake and body weight gain in rats. He then attended Florida State University in Tallahassee, FL for graduate training with Dr. Lisa Eckel in August 2011. In August 2014, Sean earned his M.S. in Psychobiology, where he studied the relative contributions of the two primary estrogen receptors, ERa and ERb, in estrogen’s effects on food intake and anxiety-like behavior in female rats. In December 2019, Sean earned his Ph.D. in Neuroscience, where he investigated the ability of the novel cannabinoid, AM11101, to increase food intake and reduce body weight loss within a preclinical rodent model of anorexia nervosa, the activity-based anorexia paradigm. To compliment his training in ingestive behavior, Sean joined the Zigman lab in July 2020 to explore the role of ghrelin in the metabolic responses to exercise and exercise endurance. Outside of the lab, he enjoys photography, hiking, building computers, cooking, and spending time with his wife and family.

Omprakash Singh, Ph.D.
Postdoctoral Research Fellow
Omprakash Singh, Ph.D.
Postdoctoral Research Fellow
Om was born and raised in Ballia district, which is the easternmost city of Uttar Pradesh, India. He earned his master’s degree in Zoology at the University of Lucknow in 2011, and then he worked as a research fellow in the Biological Rhythm Research Laboratory, University of Lucknow, where he studied the regulation of circadian and seasonal responses in birds. In July 2013, Om joined Dr. Praful Singru’s lab at NISER-Bhubaneswar, India, and completed his Ph.D. degree in May 2020. In the Singru lab, he worked on two projects: (i) determining the significance of non-hypophysiotropic TRH in controlling food intake and energy balance in the mammalian brain and (ii) determining how the TRHergic neural circuitry is organized in the brain of non-mammalian vertebrates. He employed a teleost, reptile, and bird as model animals to trace the evolution of TRHergic modulation of energy balance. Om joined the Zigman lab in October 2020. In the lab, Om is using stereotaxic surgery, neuronal tract tracing, immunofluorescence, ICV drug administration, and behavioral analysis of mice to understand the role of ghrelin in the metabolic responses to exercise and exercise endurance. Outside the lab, Om enjoys traveling, exploring nature, and playing + watching cricket.

Research Assistant
Corine Richard, M.S.
Research Assistant
Corine was born and raised in Atlanta, GA. She obtained her B.S. in Biological Science in 2015 from Georgia Southern University and her M.S. in Neurobiology and Human Behavior in 2017 from Georgia State University in Atlanta, GA. Her M.S. thesis research includes having studied how various genetic factors (including a variety of common and rare genetic variants) and environmental factors interact as potential causes for onset of Schizophrenia. She also completed neuroendocrinological research on the effect of sex difference on stress hormone levels in juvenile convict cichlids (Amatitlania nigrofasciata) while at Georgia State University. She relocated to Dallas, TX in 2019 and in October 2020, joined Dr. Zigman’s lab.
References:
2018: Satiety behavior is regulated by ASI/ASH reciprocal antagonism. Davis KC, Choi YI, Kim J, You YJ. Sci Rep. 2018 May 2;8(1):6918. doi: 10.1038/s41598-018-24943-6.
2016: Fat Metabolism Regulates Satiety Behavior in C. elegans. Hyun M, Davis K, Lee I, Kim J, Dumur C, You YJ. Sci Rep. 2016 Apr 21;6:24841.
2015: An opioid-like system regulating feeding behavior in C. elegans. Cheong MC, Artyukhin AB, You YJ, Avery L. Elife. 2015 Apr 21;4.
2013: ASI regulates satiety quiescence in C. elegans. Gallagher T, Kim J, Oldenbroek M, Kerr R, You YJ. J Neurosci. 2013 Jun 5;33(23):9716-24
2008: Insulin, cGMP, and TGF-beta signals regulate food intake and quiescence in C. elegans: a model for satiety. You YJ, Kim J, Raizen DM, Avery L. Cell Metab. 2008 Mar;7(3):249-57
2008: Lethargus is a Caenorhabditis elegans sleep-like state. Raizen DM, Zimmerman JE, Maycock MH, Ta UD, You YJ, Sundaram MV, Pack AI. Nature. 2008 Jan 31;451(7178):569-72.
2006: Starvation activates MAP kinase through the muscarinic acetylcholine pathway in Caenorhabditis elegans pharynx. You YJ, Kim J, Cobb M, Avery L. Cell Metab. 2006 Apr;3(4):237-45
- Postdoctoral Fellows/Instructors
- Graduate Students
- Research Technicians & Research Associates
- Summer Students
Ichiro Sakata, Ph.D. – Postdoctoral Research Fellow 2007 – 2010 – Ichiro is currently Associate Professor in charge of his own lab at Saitama University in Japan.
Mario Perello, Ph.D. – Postdoctoral Research Fellow 2/2008 – 3/2010 – Mario is currently an associate investigator in charge of his own lab at the Laboratory of Neurophysiology, Multidisciplinary Institute of Cell Biology (IMBICE) in La Plata, Argentina
Jen-Chieh (Jay) Chuang, Ph.D. – Postdoctoral Research Fellow 8/2008 – 5/2011 – Jay is currently working as a Biomarker Scientist at Gilead in the San Francisco Bay Area.
Paul K. Piper Jr, M.D. – Postdoctoral Research Fellow/Clinical Fellow in Diabetes, Endocrinology, and Metabolism, 2009-2011 – Paul is currently in private practice as an endocrinologist in The Woodlands, TX.
Daniela Pereira Derderian, Ph.D., Pharm.D. – Postdoctoral Research Fellow 5/2010 – 7/2012 –Daniela is currently an assistant professor in biological sciences at the School of Math and Science at Wayland Baptist University in Plainview, TX.
Won-Mee Park, Ph.D. – Postdoctoral Research Fellow – 2011–2012 – Won-Mee is currently enjoying her role as a new mom.
Qian Wang, Ph.D. – Postdoctoral Research Fellow – 1/2012–12/2013 – Qian is currently working as a Data Scientist at AIG.
Aki Uchida, Ph.D. – Postdoctoral Research Fellow – 1/2012–3/2014 – Aki is currently a Senior Project Manager at Takeda.
Sun-Hyun Park, Ph.D. – Postdoctoral Research Fellow – 10/2015–9/2016 – Sun–Hyun has moved back to South Korea where he is pursuing a science-related business venture.
Emily Bruggeman, M.S., Ph.D. – Postdoctoral Research Fellow –7/2016 – 7/2017 – Emily is currently working as a neuroscientist at Emory University School of Medicine.
Shota Takemi, Ph.D. – Postdoctoral Research Fellow – 6/2019 – 12/2019 – Shota is an Assistant Professor at Saitama University, Japan
Bharath Mani, D.V.M., Ph.D. – Postdoctoral Research Fellow, Instructor – 5/2012 – 5/2020 – Bharath is currently working as a scientist at Novo-Nordisk in Indianapolis, IN
Angela Walker – Ph.D. student in the Neuroscience Graduate Program, 1/11–4/14 – Angela successfully defended her thesis entitled “From the ghrelin cell to ghrelin action: Assessing ghrelin’s influence on mood, cue-potentiated feeding, and ghrelin cell physiology.” She is performing basic and clinical research related to autism at UT Southwestern.
Siegfried Meier – Research Assistant II, 3/09–3/10
Sherry Rovinsky – Research Assistant, 2007-2009. Graduate Student, 2010–2011.
Chelsea Migura – Research Assistant I, 3/2010–7/2012
Brittany Mason, Ph.D. – Senior Research Associate, 6/2012–7/2013
Sydney Lawrence – Research Assistant, Summer 2013, Summer 2014, Summer 2015, Summer 2016
Connor Lawrence – Research Assistant, Summer 2015, Summer 2016
Jakub Woloszyn – 2008 UTSW Medical Student, Summer Research
Neha Chaudhary – 2009 UTSW Medical Student, Summer Research
Gregory Wallingford, Jr. – 2009 Summer Undergraduate Research Fellowship student
Anna Kuperman – 2010 Summer Undergraduate Research Fellowship student
Shloka Raghavan – 2011 Summer Undergraduate Research Fellowship student
Carolyn Chakuroff – 2012 Summer Undergraduate Research Fellowship student
Imikomobong (Micky) Ibia – 2012 MSTP Summer Undergraduate Research Fellowship student
Christina Mosher –2013 UTSW Medical Student, Summer Research
Madhu Karamsetty – 2013 Summer Undergraduate Research Fellowship student
Nicole Huang – 2014 Summer Undergraduate Research Fellowship student
Rachel Hodge – 2015 Summer Undergraduate Research Fellow
Hannah Viroslav –2016 UTSW Medical Student, Summer Research
Henry Roseman –2017 Undergraduate student, Summer Research
Anna Patterson –2019 Amgen scholar
Avi Burstein –2019, 2020 High School student, Summer Research
Available Positions
Available Student, Postdoctoral fellow, and Research Technician Positions
The Zigman lab is seeking motivated, hard-working, and imaginative student, postdoc, and technician researchers to join our team.
The Zigman lab investigates the neuronal/hormonal basis for eating, body weight, and blood glucose control with the goal of designing new methods to treat obesity, diabetes, hypoglycemia,
cachexia/anorexia, and associated disorders of mood. Most of our studies focus on the hormone ghrelin, which is made primarily by a distinct group of cells in the stomach, the ghrelin receptor (GHSR), which is highly expressed in the brain and pancreatic islets, and LEAP2, which decreases activity of the ghrelin receptor. Our past and ongoing studies have investigated how ghrelin and ghrelin receptors influence eating, blood glucose, body weight, and mood in the settings of high fat diet exposure, caloric restriction, exercise, chronic stress, weight loss surgery, insulin administration, and Prader-Willi Syndrome. We also study the mechanisms of ghrelin secretion. Newer projects investigate novel proteins involved in the body’s responses to obesity and caloric restriction.
Several graduate student and postdoctoral positions are available in the Zigman lab. There are also Research Assistant/Research Associate positions available. Projects will take advantage of our one-of-a-kind collection of transgenic mouse models targeting the ghrelin system, other endocrine cells, and neurons. These models allow us to characterize and manipulate gene expression and modulate cellular activity in specific cells and neurons of interest. Methods include mouse behavioral protocols, CRISPR-Cas9, transcriptomics, tissue clearing, cell culture, histology, glucose clamp techniques, stereotaxic brain surgery, chemogenetics, optogenetics, and in vivo fiber photometry, to name a few.
Candidates for the graduate student positions can come from any UT Southwestern Ph.D. Program, and most usually are from the Neuroscience Graduate Program or the Genetics Development and Disease Graduate Program. Students most often are also affiliated with the Molecular Metabolism and Metabolic Diseases (3MD) Track (chaired by Dr. Zigman). We would love to have you and your friends rotate through the lab!
Candidates for the postdoctoral positions must hold a Ph.D. and/or M.D. degree.
Graduate student and postdoctoral fellow candidates should be self-motivated and are expected to contribute substantively to the design, implementation, interpretation and reporting of their investigational studies. Prior experience with genetically-engineered mouse models and related breeding strategies, mouse behavioral studies, stereotaxic brain surgery and other neuroanatomical techniques such as chemogenetics and optogenetics, histology, cell culture, and/or bioinformatics leading to publication in peer-reviewed journals is recommended but not required.
Interested individuals should send a CV, statement of interests, and a list of three references to Jeffrey Zigman, M.D., Ph.D.
UT Southwestern Medical Center is an Affirmative Action/Equal Opportunity Employer. Women, minorities, veterans and individuals with disabilities are encouraged to apply.

Dr. Elmquist appointed Vice Chair of Research for Internal Medicine
Joel Elmquist, D.V.M., Ph.D., Professor of Internal Medicine, has been

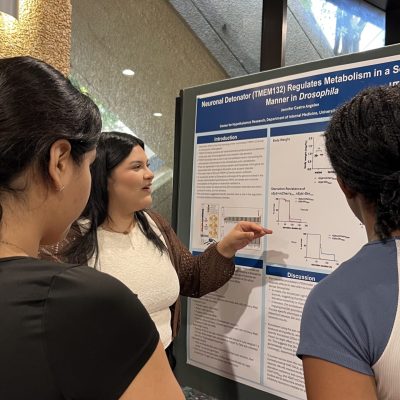
Neuronal E93 Regulates Metabolic Homeostasis
Cecilia Yip1, Steven Wyler1, Syann Lee1, Adrian Rothenfluh2, Young-Jai You1,

Insulin receptors directly regulate feeding-induced dips in plasma ghrelin
A recent study by the Zigman lab, spearheaded by Dr.